Publications
Our scientists regularly release their work to benefit the field. Here is a selection of our contributions.
Underpinning everything in our Platform is a commitment to robust information and software systems - they enable scalable workflow-integrated AI, and reliability, reproducibility and correctness, ensuring we advance molecules with the best chances of helping patients.
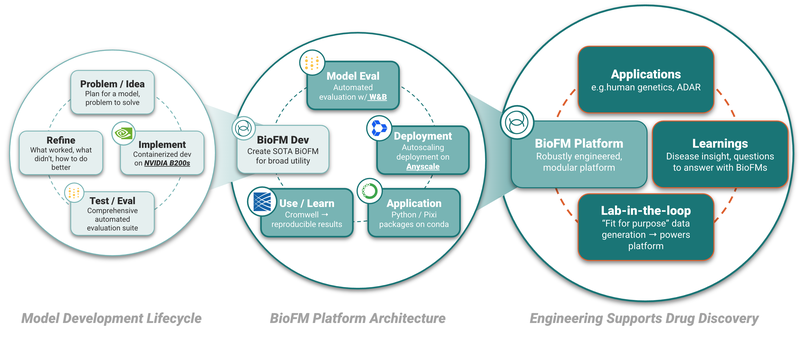

BigRNA is a proprietary, powerful BioFM that was trained on over a trillion signals derived from high-throughput sequencing datasets and has distilled the core principles of RNA biology. This allows it to serve as a versatile model for numerous downstream applications, such as predicting the effects of genetic variants, fine-tuning for predicting on- and off-target effects of genetic medicines, and generating rich sequence embeddings for other models used in target biology discovery and molecular design. Our BigRNA workflows include Weights and Biases integration, sophisticated model evaluation frameworks and provide meaningful scalar outputs while enabling fine tuning on new datasets.
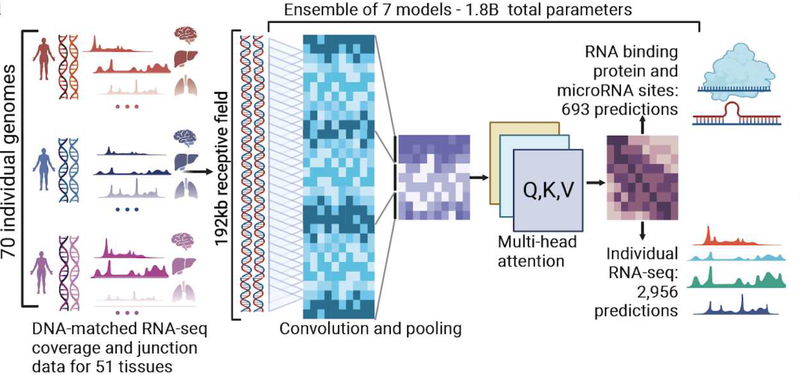
Since its initial development in 2021, BigRNA has evolved significantly. Key enhancements include a shift to higher prediction resolution (from 128-bp to 1-bp), the integration of new training datasets (e.g. relevant to disease contexts), and architectural improvements enabling higher predictive power and efficiency.

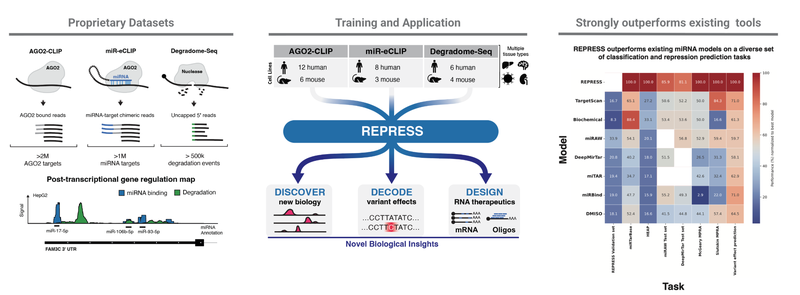
REPRESS is a deep learning foundation model that predicts cell-type-specific microRNA (miRNA) binding and mRNA degradation directly from RNA sequence. It has been shown to reveal biology missed by other state-of-the-art methods, such as identifying repressive non-canonical miRNA target sites and decoding the regulatory effects of sequence context and miRNA binding site multiplicity.
REPRESS outperforms other advanced methods and neural architectures on a comprehensive suite of orthogonal tasks, including identifying genetic variants that affect miRNA binding, predicting out-of-distribution data from massively parallel reporter assays, and predicting canonical and non-canonical miRNA mediated repression.

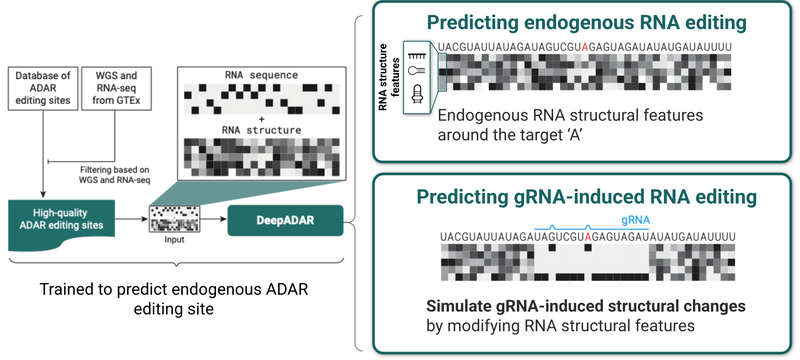
DeepADAR can design guide RNAs (gRNAs) to induce ADAR-mediated editing across various trinucleotide contexts. The base DeepADAR model is trained to predict endogenous ADAR editing based on local sequence and structure around candidate editing sites. By observing endogenous editing at 16 million target sites, as well as a multitude of other sites where editing does not occur, the base DeepADAR learned the subtle sequence and structural features that direct ADAR to edit specific sites. Using screening data from a custom dataset of synthetic gRNAs, we fine-tuned this base model to predict gRNA-driven editing, based on sequence and structural features created by gRNAs.
Deep Genomics views data generation and experimental validation as crucial components of our BioFM Platform. To develop fit-for-purpose training and validation datasets plus datasets for therapeutic programs, we operate two experimental facilities, with over 10,000 square feet of lab space between Toronto, Ontario and Cambridge, Massachusetts. In both locations, our experimental scientists work hand in hand with their machine learning and computational counterparts to design experiments and workflows that accelerate our mission.

Our scientists regularly release their work to benefit the field. Here is a selection of our contributions.