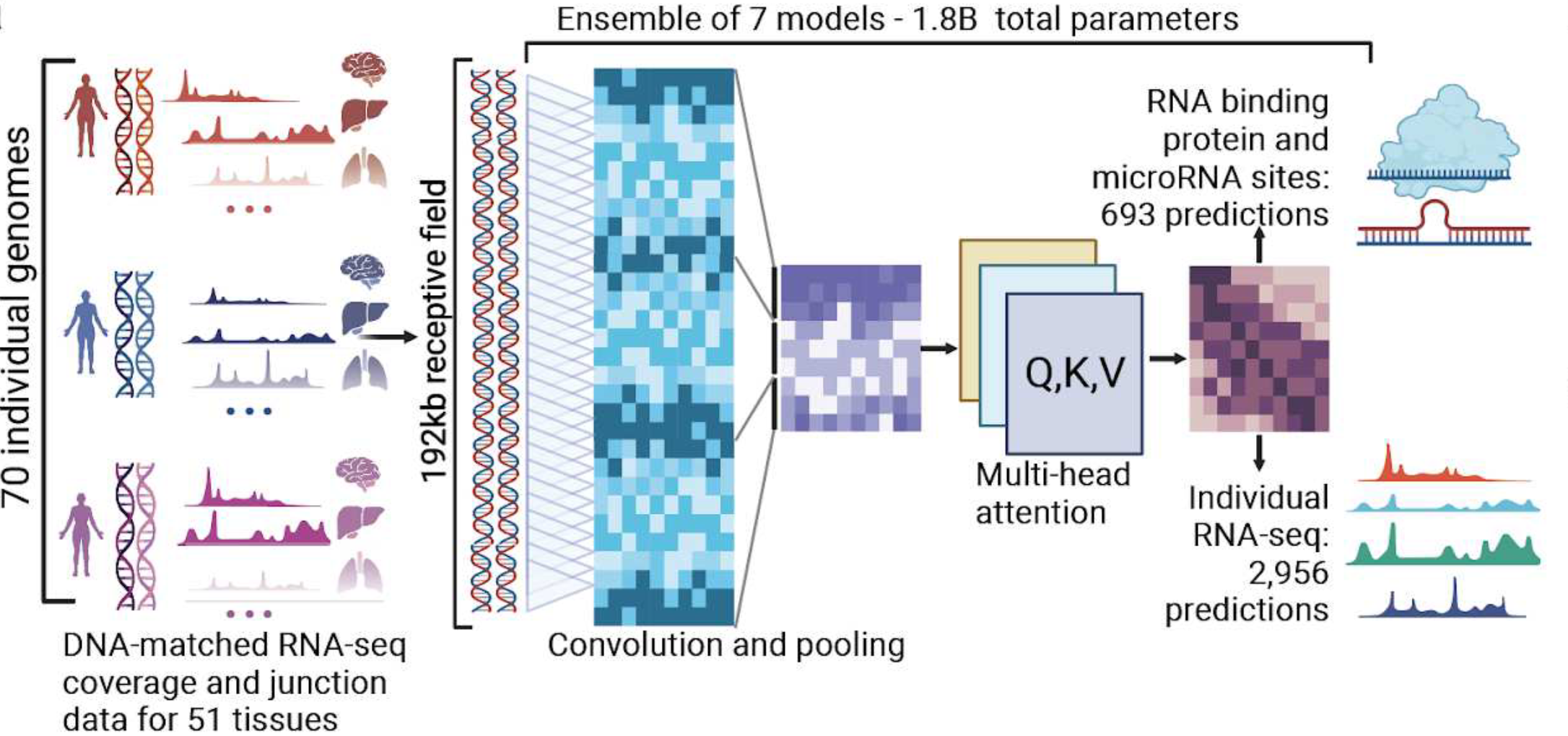
Company’s unique AI foundation model BigRNA is the first transformer neural network for the discovery of RNA biology and therapeutics
Represents a new generation of deep learning AI that can be applied to a range of different RNA therapeutic discovery tasks
Comprised of nearly two billion tunable parameters and trained on thousands of datasets comprising one trillion genomic signals
TORONTO--(BUSINESS WIRE)--September 27, 2023 Deep Genomics, a leading AI drug development company focused on decoding biology to program life-changing medicines, announced today the release of the manuscript, “An RNA foundation model enables discovery of disease mechanisms and candidate therapeutics” introducing the company’s AI foundation model, BigRNA. As outlined in the manuscript, BigRNA accurately predicts the tissue-specific regulatory mechanisms of RNA expression and the binding sites of proteins and microRNAs, plus the effects of variants and candidate therapeutics. Unlike existing approaches that are tailored to a single task, foundation models generate wide and general outputs, meaning BigRNA can uniquely discover new biological mechanisms and RNA therapeutic candidates that would not be found using traditional approaches.
“Building machine learning models that can predict gene expression from DNA sequence has been a long-standing research goal, and one that has seen significant strides owing to recent advancements in deep learning. Deep Genomics has shown that our unique AI foundation model, BigRNA, can utilize DNA sequences to accurately discover the effects of non-coding, missense and synonymous variants on tissue-specific gene regulation, identify new mechanisms of RNA biology, and design RNA therapeutic candidates,” said Brendan Frey, Ph.D., F.R.S.C., founder and chief innovation officer at Deep Genomics. “In a comprehensive study of 14 genes, BigRNA consistently designed highly effective steric blocking oligonucleotides that act in a tissue-specific manner, including for genes involved in Wilson disease and spinal muscular atrophy. We believe that BigRNA and deep learning systems like it have the potential to transform the field of RNA therapeutics.”
“Kudos to Brendan and the team at Deep Genomics for their remarkable research on developing the first foundation model for RNA therapeutics. The engineering effort required to build systems like this is extraordinary and Deep Genomics has achieved it. I'm excited to see the latest advancements in machine learning make their way to meaningful applications,” said Yann LeCun, Ph.D., chief AI scientist at Facebook, Professor at New York University and member of the scientific advisory board at Deep Genomics.
BigRNA learns from paired genotype and high resolution RNA expression data from many individuals, and can also be applied in a range of downstream tasks such as predicting RNA-binding protein (RBP) and microRNA binding sites. BigRNA can also help design different types of RNA based therapeutics, including steric blocking oligonucleotides (SBOs). Without any additional training, BigRNA accurately identifies compounds that induce a targeted splicing change, and recovers known approved SBO therapies with high specificity. The ability of BigRNA to understand regulatory mechanisms also allows it to design SBOs that block predicted inhibitory regions to increase the expression of a disease gene. Publication of this manuscript and presentation of this data is expected at future scientific meetings and journals.
Key Outcomes Highlighted in the Manuscript:
Advances in deep learning and molecular biology have introduced the possibility of building systems that can discover the mechanisms regulating RNA processing, predict the effects of genetic variants, and design therapeutic molecules that restore RNA and protein. These models could revolutionize drug discovery by pinpointing how pathogenic genetic variants alter gene expression and gene processing, and by designing customized drug candidates to counteract these effects.
Currently, most efforts have focused on predicting data that measures overall gene expression levels, which are not suited to predicting regulatory interventions, for example, specific transcriptional perturbations on splicing or polyadenylation. In contrast, BigRNA is trained to predict RNA expression at sub-gene resolution.
BigRNA can be used to design SBOs that alter RNA processing and reverse the effect of a pathogenic variant or increase or decrease expression. BigRNA accurately predicted the effects of SBOs on increasing the expression for all of 4 genes, and on splicing for all of 18 exons across 14 genes, including those involved in Wilson disease and spinal muscular atrophy. When we compared BigRNA to the current state of the art in deep learning technology for predicting the binding sites of 153 proteins and micro-RNAs, BigRNA performed substantially better in 152 of cases, and tied the state of the art in only one case.
When used for target discovery by evaluating genetic variants, BigRNA was able to accurately identify pathogenic variants along with their molecular mechanisms. It correctly predicted that a pathogenic variant in the 3’UTR of NAA10 alters the polyadenylation site (syndromic X-linked microphthalmia), that a pathogenic variant in KCNH2 causes intron retention (long QT syndrome), and that the M645R variant in ATP7B causes exon skipping (Wilson disease). BigRNA is competitive on classification metrics, e.g., for a database of 3’UTR variants, BigRNA detected 83% of known pathogenic variants at a false positive rate of 5%, substantially outperforming an RNA stability model that achieved a detection rate of 50%.
About BigRNA
BigRNA is the first foundation model for RNA that has been shown to be effective for a range of drug discovery tasks, including predicting the effects of patient variants on RNA processing mechanisms, discovering novel regulatory RNA biology, and designing steric blocking oligonucleotides that alter splicing or increase gene expression. BigRNA is a transformer-based deep learning system trained on thousands of RNA-seq datasets, comprising over a trillion genomic signals. It can be used to discover the effects of non-coding, missense, and synonymous variants and design therapeutic candidates. Provided with unannotated DNA or pre-mRNA sequence as input, BigRNA can predict RNA expression, splicing, the binding sites of microRNAs and RNA binding proteins, and the effects of RNA therapeutic candidates, across a range of specific tissues and genetic backgrounds.
About AI Foundation Models
AI foundation models have emerged in the last three years and are quickly transforming entire industries. They are very large machine learning models, trained on vast quantities of data at scale, such that they can be adapted to a wide range of downstream tasks. A traditional AI model could be very good at one task, such as predicting molecule-target interaction, whereas a foundation model has learned fundamental aspects that benefit many tasks, such as target identification, predicting molecule-target interactions and designing molecules. Their value is based on their limitless potential.
About Deep Genomics
Deep Genomics is a biopharmaceutical company that aims to revolutionize drug development by leveraging expertise in artificial intelligence (AI) to decode RNA biology. Our proprietary platform, the AI Workbench, enables us to decode the enormous complexity of RNA biology to find novel targets, mechanisms, and molecules that are not accessible through traditional methods. We use this advanced technology to develop steric-blocking oligonucleotides (SBOs) that achieve expression increase for the treatment of genetic disease. Founded in 2015, our multidisciplinary team includes expertise in a diverse range of disciplines including those found in a traditional drug company, as well as machine learning, laboratory automation, and software engineering. Deep Genomics is located in Toronto, Ontario and Cambridge, Massachusetts. For more information, visit: www.deepgenomics.com and follow us on LinkedIn and Twitter.
Media Contact:
Deep Genomics
press@deepgenomics.com